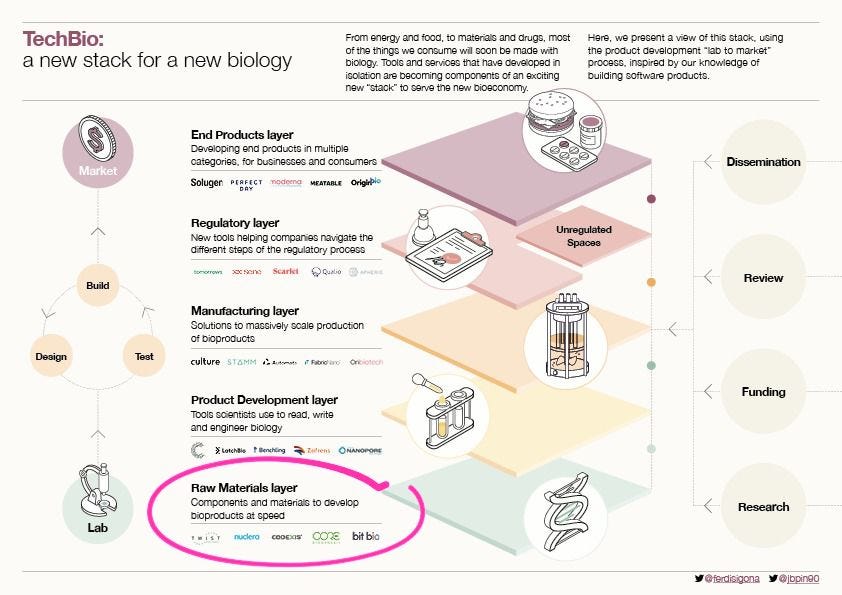
Whilst AI tools like AlphaFold have made a splash on scientific and popular media covers, it is a mostly quiet revolution in our ability to manipulate biology’s building blocks that, we believe, is behind the emergence of biology as an engineering discipline. From the stone age to the digital age, our use of materials has played a key role in the evolution of the global economy. The same is true as we enter the age of biology. In this post, we explore our fast-developing ability to write biology’s code, keeping in line with our analogy: the raw materials layer of the TechBio Stack.
As the central dogma of molecular biology goes, the genetic flow of information is: DNA encodes RNA, which in turn encodes proteins. If DNA is the code script, RNA is the compiler and proteins the executable programs which perform any function within biological systems. Unlike the hardware and software we’ve built in computer science, however, we’ve had to reverse engineer biology by first observing complex programs in action. We’ve since learned to read (sequence), write (synthesise) and edit (CRISPR) DNA, accelerating and massively parallelising these two processes is making biology programmable.
DNA
DNA is, in essence, information: a bacterium typically contains 750 kB of DNA, stored in a single (long) molecule, while human DNA is stored in 46 molecules (chromosomes), and contains around 750 MB. Putting that into more accessible terms, if we took our raw DNA data and if we could play it back as uncompressed CD quality audio, our genome would be roughly one hour of sound (the average run time of a full-length music album), the DNA of the smallest bacterium would be a 1.5 second sound clip, and the smallest virus would just emit a tiny blip two milliseconds long. Our ability to read DNA has evolved stupendously over the past 15 years. From old-school Sanger sequencing, which enabled us to read a single, short DNA snippet at a time, we moved on to next-generation sequencing, which works through millions of fragments simultaneously. Sequencing chemistry has been miniaturised and put on silicon chips, making the process fast, cheap and readily available.
If we are to programme biology, it’s important we don’t just read its code, but write it as well. Our ability to do this has also evolved exponentially. With PCR, we are able to copy” the code with amazing accuracy: an error rate of almost 1/2,000,000. Why is copying DNA useful? PCR is used in many research labs, and it also has practical applications in forensics, genetic testing, and diagnostics. By using PCR to “amplify” a specific genetic sequence in a sample we are able to more easily detect its presence, which has become commonplace for COVID testing but is also used for detecting other pathogens or even specific genetic diseases.
The ability to copy DNA is actually a critical piece of infrastructure for manufacturing next generation therapeutics like many viral and mRNA therapies today. Plasmids are small fragments of DNA, which can be engineered to introduce gene(s) of interest into cells. There are not many good ways to rapidly scale DNA plasmid production and as the cell and gene therapy field evolves, demand for longer and GMP-grade DNA will continue to rise. Some startups are tackling this issue, such as Touchlight, a UK based company pioneering a way to make plasmids easy.
With gene synthesis, we are able to “write” the code from scratch, but we are - again - still very early: we can’t even synthesise a whole human genome yet. Long DNA snippets - called oligos - need to be synthesised and joined together to form genes. Ten years ago, DNA synthesis was a slow and difficult chemical process. The four bases (A, T, C, and G) that make up DNA were used as reagents, and they were pipetted onto a plastic plate with 96 pits, or wells, each of which held about 50 microliters, or one eyedropper drop of liquid.
Today, companies like Twist Bioscience have massively scaled this process by moving it onto silicon chips able to hold millions of tiny, nL scale wells, and replacing pipetting with an inkjet printer. The result of this is a 10x decrease in the cost of producing DNA, which has led to a massive democratisation of the technology. Today, customers can visit the Twist website, upload their DNA sequence, pay, and get DNA delivered to their lab in a few days, to be used for producing everything from drugs, to food flavourings, to industrial products.
Twist’s 2021 IPO prospectus shines a light on the exciting future of this technology:
The ability to design DNA and engineer biology is creating advances and benefits for a broad and growing range of applications for synthetic DNA and synthetic DNA-based products across multiple industries, including:
healthcare for the discovery and production of new vaccines, therapeutics and molecular diagnostics;
chemicals/materials for cost-effective and sustainable production of new and existing specialty chemicals and materials, such as spider silk, nylon, rubber, fragrances food flavours and food additives;
food/agriculture for more effective and sustainable crop production;
academic research for a broad range of applications; and
technology for potential use as an alternative long-term data storage medium.
Continued miniaturisation and massive parallelisation enabled by advances in silicon technology and microfluidics will lead to further reductions in the cost and improvements in the DNA synthesis lengths that are achievable today. Companies like Elegen sit on this new frontier.
But, just like in DNA sequencing, alternative technologies also have a role to play. An example of this is enzyme-based modes of DNA synthesis, which could potentially improve synthesized DNA length and quality. As a general rule, enzymes can perform reactions reliably in aqueous conditions in which DNA is stable. This enzymatic approach has been proposed for decades and has been under development by research groups and companies like DNA Script in recent years.
The final piece in our DNA writing toolbox (from copying, to de novo synthesis, to editing) is CRISPR. With CRISPR (awarded the Nobel Prize in Chemistry in 2020), we are now able to not just copy paste, but “edit” pre-existing long strands of DNA code. It acts as a precise pair of molecular scissors that can cut a target DNA sequence, directed by a customizable guide. CRISPR - alongside other, less famous “edit” tools in development - has the potential to disrupt many aspects of life from gene editing therapies, to disease resistant crops or advanced biofuels so it deserves its own post.
RNA
DNA and proteins have taken up most of the public’s attention over the last decades given how much the biotech industry has relied on the commercialisation of both. RNA has only come to the forefront because of the covid 19 pandemic and the new mRNA vaccines developed by BioNTech and Moderna. However, neither of those companies specialises in manufacturing RNA, only designing the code snippet which will be injected into our bodies.
First of all, it’s important to note the difference between DNA and RNA when diving into how it’s made. RNA is typically single stranded (incl the 3 main types involved in protein synthesis mRNA, tRNA and rRNA) and composed of 4 bases with one difference vs DNA (AUCG vs ATCG). Given the similarities, it’s natural that RNA is manufactured similarly to DNA. For short strands it can be chemically synthesized, swapping the step which includes T for U, which could potentially be scaled up to industrial scale (as well as enzymatic approaches). For longer and industrial scale use (eg. mRNA vaccines) the main methodology used is called In Vitro Transcription which is a complex biological process (see below). Many companies are focused on building novel RNA design or manipulation platforms while only a few are trying to make manufacturing more efficient. EnPlusOne is a new company spinning out of the Wyss Institute for Biologically Inspired Engineering at Harvard University working on a novel enzymatic RNA synthesis platform.
Proteins
If DNA/RNA is the code, Proteins are the compute programs. Proteins are the basic functional and structural units of pretty much all living matter around us; they play a fundamental role both in traditional biotechnology focused on healthcare, and in the more recent shift to the bioeconomy. Drug discovery starts at the protein level: to discover or design a new drug a target must be found first, these are typically proteins. They can also be used as drugs to treat disorders including diabetes, cancer and arthritis, produced as the building blocks of alternative foods (think synthetic meat) and, as enzymes, play a key role in catalyzing chemical and biological reactions to replace more traditional manufacturing systems (e.g. in materials).
The journey from gene to protein is complex and tightly controlled within each cell. It consists of two major steps: transcription and translation. Together, transcription and translation are known as gene expression. Replicating this process in the lab has traditionally involved genetically engineering microbes or other cells to produce the desired protein: this method is called recombinant expression. The first successful attempt made biotech history and was achieved by Genentech, who produced insulin. Since then, scientists have worked hard to reduce the time and complexity of the process and are even engineering proteins for better properties. This renewed focus on proteins in healthcare and beyond has meant increased investment.
Some players have focused on improving recombinant expression. According to Alexandre Reeber, "the speed of adoption of the bioeconomy will be strongly dependent on our ability to produce proteins in a sustainable and cost-efficient manner, at scale." Alexandre is the co-founder and CEO of Core Biogenesis, a French company that is focusing on a specific type of proteins, growth factors. The company specialises in using plants as the host organism, and then uses its expertise to tackle the key cost drivers: low production yield due to gene silencing (a natural limitation to gene expression) and the difficult extraction process.
Other players have focused on fermentation: an example is Perfect Day, an American company that is focusing on foods. Cultured dairy is created by taking cow's milk DNA and adding it to a microorganism like yeast to create dairy proteins, whey and casein, via fermentation.
Other players yet have focused on cell-free synthesis: an example is FabricNano, a British company focusing on enzyme immobilization, which has the potential to improve bioprocess economics in a range of industries, from manufacturing to pharmaceuticals and commodity chemicals.
Finally, desktop bioprinters have started to appear recently, which use the miniaturisation and parallelisation principles seen in DNA synthesis to massively scale up protein synthesis obtained via cell-free biology. An example is Nuclera, a UK/US company whose bioprinter combines cell-free protein synthesis with digital microfluidics to enable rapid protein prototyping: the user inputs a DNA sequence into the cartridge, which performs a series of tests at nanolitre scale to optimise protein yield and purifiability. The proteins are then produced at microlitre scale using “bio-inks” and purified using “digital microfluidics” (enables digital control of hundreds to thousands of droplets). All the user has to do is pipetting.
Cells
The final piece of the biological building blocks are cells. Cells are regarded as the basic building blocks of life. The smallest entity generally considered to be living is a single cell, and all life forms are either uni- or multicellular organisms. The bottom-up construction of an artificial cell that can be truly considered “alive” is still an ambitious goal. Current state-of-the-art artificial cells mimic the hierarchical architecture of eukaryotic and (mostly) prokaryotic cells and tissue, and attempt to reproduce most of the basic functions, but are still not good enough to replace actual cells.
Today, most cells used in labs or the clinic are derived from actual cells. Stem cells have gotten a lot of attention over the last decade, especially Induced Pluripotent Stem Cells (iPSC). These are mostly derived from skin or blood cells that have been turned back into an state that resembles their embryonic lives. By then reprogramming them to other states or types of cells an unlimited source of any type of human cell needed for therapeutic purposes is unlocked. Bit.bio in the UK has developed the opti-ox platform to efficiently reprogram human cells.
Final Thoughts
Materials have defined human’s capabilities since the dawn of civilization, leading to all previous revolutions, from agriculture, to industry and informatics. Now, our ability to fundamentally recreate biology is behind the life sciences and biology revolution. DNA, RNA and protein synthesis have evolved exponentially over the last decades, fueled by an unparalleled development of market demand for next generation therapies and products. In the future, we are certain we will see more complex biological systems being built from scratch. As we dive deeper into the stack we hope we can continue to unpack this constantly evolving field.